ARTICLE BY MR. EZRA SALAMAT
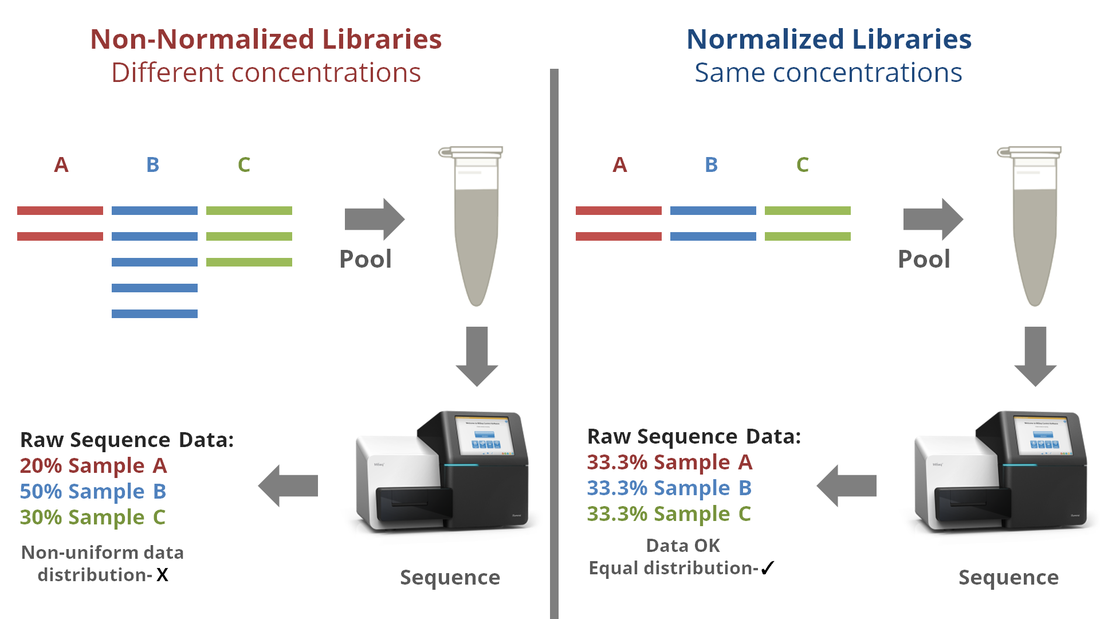
STEP 6: LIBRARY NORMALIZATION
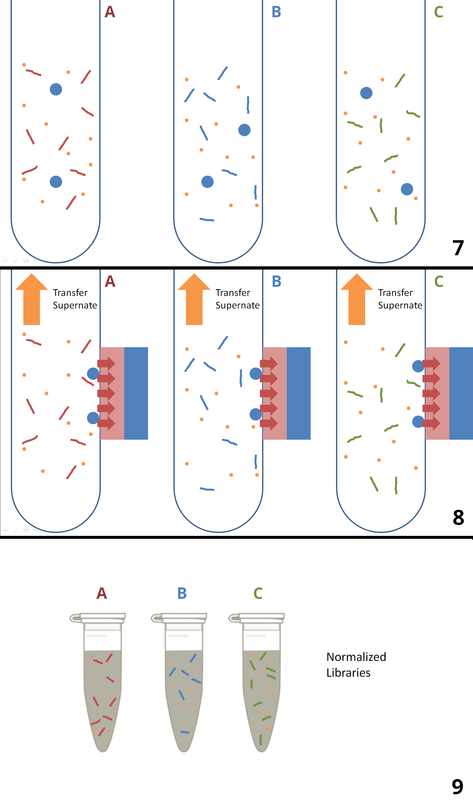
Normalization can be done by using magnetic beads (similar to the one used in index PCR cleanup), or by dilution with the aid of a fluorometer. In this chapter, we will discuss only the bead-based normalization. Normalization starts with cleaned Index-PCR products, each of which has a different library concentration (1). Am equal concentration of SPRI (magnetic) bead is added to each sample (2). As each bead binds an identical number of DNA strands, an identical number of DNA libraries bind to the added beads across all samples (3). A strong magnetic field is then applied, causing the beads (with bound libraries) to stick to the side of the tube (4). The supernates containing excess libraries are then washed out, leaving the beads with bound libraries behind. (5). Elution buffer is then added to the tubes (6). The elution buffer causes the DNA libraries to be released from the magnetic beads (7). The tubes are exposed again to a strong magnetic field, which causes the beads (now w/ no DNA libraries bound) to stick to the side of the tubes (8). The supernates, now containing equal concentrations of DNA libraries, are then transferred to separate tubes (9). STEP 7: POOLING AND Denaturation
The libraries must then be denatured, to allow the single strands to bind to the Illumina flow cell that contains immobilized oligos. These immobilized oligos are complementary to the P5 and P7 sequences introduced during Index PCR. To do this, the pooled libraries are diluted then heated to 98 °C, separating the double strands into two. The tube is then cold-shocked to 5 °C. The rapid cooling prevents the denatured libraries from reannealing. An internal control called Phi-X is also added. (see figure below). The Phi-X in the Illumina sequencing kit is an adapter-ligated library of the Phi-X 174 (or ΦX174) bacteriophage genome. Phi-X 174 is is a single-stranded DNA virus, of which the first DNA-based genome was successfully sequenced. This was done using a technique (involving DNA polymerization coupled with slab gel electrophoresis) performed by Fred Sanger and his team in 1974. The genome of Phi-X is small, balanced in nucleotide concentration, and well characterized, and as such is used as a reliable internal control for troubleshooting and evaluating sequencing runs. STEP 8: Sample Loading and SequencingAfter the library has been prepared, it's time to load it to the sequencer, along with the necessary components. Shown in the figure below is the MiSeq Sequencing kit. The Miseq Reagent Cartridge (1) contains all reagents (modified dNTPs, dye terminator remover, buffers, etc) for sequencing in one convenient package. It is stored frozen, and must be thawed on water then mixed before use. The Phi-X internal control (2) must be denatured and normalized, as was used earlier during the denaturation step. The MiSeq Flow Cell (3) is stored in buffer. It must be washed with distilled water, then dried before use. The Incorporation Buffer (4) is used as is. The pooled and Denatured Libraries w/ Phi-X are then loaded into the MiSeq Reagent Cartridge, as shown in the figure below. The Cartridge and Incorporation Buffer are then loaded into the instrument, as shown. The Miseq Flow cell, once washed and dried, is also loaded into the machine and clamped down under the holder assembly.
After the run, the machine will produce an output of up to 15Gb worth of data, amounting to up to 25 million reads. This is still raw data comprising of ~300bp long reads, each with an identifier tag and quality score line. These must be processed via software to generate data useful for scientists. This concludes the wetlab portion of the Illumina DNA Sequencing workflow. In the next chapter, we'll take a look at the in silico part of DNA sequencing, where we use computers and software to assemble and analyze the raw data output of the next-generation sequencers. NEXT: PART IV |
| Interested in other Illumina Workflows, Platforms, and Applications> You can head over to the Illumina Youtube channel at: https://www.youtube.com/user/IlluminaInc |
References:
- Heather JM, Chain B. The sequence of sequencers: the history of sequencing DNA. Genomics. 2016;107:1–8. doi: 10.1016/j.ygeno.2015.11.003.
- Nextera XT DNA Library Preparation Kit Data Sheet. Retrieved from: https://www.illumina.com/products/by-type/sequencing-kits/library-prep-kits/nextera-xt-dna.html
- Illumina Miseq Plaform. Retrieved from: https://www.illumina.com/systems/sequencing-platforms/miseq.html
2 Comments
Peter Bell
8/26/2021 09:53:02 am
Nice overview. I think Fred Sanger in 1977 used a slab gel instead of capillary electrophoresis. Might be wrong about that, but having done a ton of Sanger and Maxam Gilbert sequencing in the 1980s, I think that slab gels were most likely used.
Reply
Ezra Salamat (Author)
8/26/2021 10:30:37 am
Yes, you are correct, and our apologies for the mistake. I have made the necessary revision. Thank you!
Reply
Leave a Reply.
Archives
April 2018
February 2018
October 2017